| [ < ] | [ > ] | [ << ] | [ Up ] | [ >> ] | [Top] | [Contents] | [Index] | [ ? ] |
21.1.4.3 Mathematical Considerations
The attempt has been made to make sparse matrices behave in exactly the same manner as there full counterparts. However, there are certain differences and especially differences with other products sparse implementations.
Firstly, the "./" and ".^" operators must be used with care. Consider what the examples
s = speye (4); a1 = s .^ 2; a2 = s .^ s; a3 = s .^ -2; a4 = s ./ 2; a5 = 2 ./ s; a6 = s ./ s; |
will give. The first example of s raised to the power of 2 causes
no problems. However s raised element-wise to itself involves a
large number of terms 0 .^ 0 which is 1. There s .^
s is a full matrix.
Likewise s .^ -2 involves terms like 0 .^ -2 which
is infinity, and so s .^ -2 is equally a full matrix.
For the "./" operator s ./ 2 has no problems, but
2 ./ s involves a large number of infinity terms as well
and is equally a full matrix. The case of s ./ s
involves terms like 0 ./ 0 which is a NaN and so this
is equally a full matrix with the zero elements of s filled with
NaN values.
The above behavior is consistent with full matrices, but is not consistent with sparse implementations in other products.
A particular problem of sparse matrices comes about due to the fact that as the zeros are not stored, the sign-bit of these zeros is equally not stored. In certain cases the sign-bit of zero is important. For example
a = 0 ./ [-1, 1; 1, -1];
b = 1 ./ a
⇒ -Inf Inf
Inf -Inf
c = 1 ./ sparse (a)
⇒ Inf Inf
Inf Inf
|
To correct this behavior would mean that zero elements with a negative sign-bit would need to be stored in the matrix to ensure that their sign-bit was respected. This is not done at this time, for reasons of efficiency, and so the user is warned that calculations where the sign-bit of zero is important must not be done using sparse matrices.
In general any function or operator used on a sparse matrix will
result in a sparse matrix with the same or a larger number of non-zero
elements than the original matrix. This is particularly true for the
important case of sparse matrix factorizations. The usual way to
address this is to reorder the matrix, such that its factorization is
sparser than the factorization of the original matrix. That is the
factorization of L * U = P * S * Q has sparser terms L
and U than the equivalent factorization L * U = S.
Several functions are available to reorder depending on the type of the matrix to be factorized. If the matrix is symmetric positive-definite, then symamd or csymamd should be used. Otherwise amd, colamd or ccolamd should be used. For completeness the reordering functions colperm and randperm are also available.
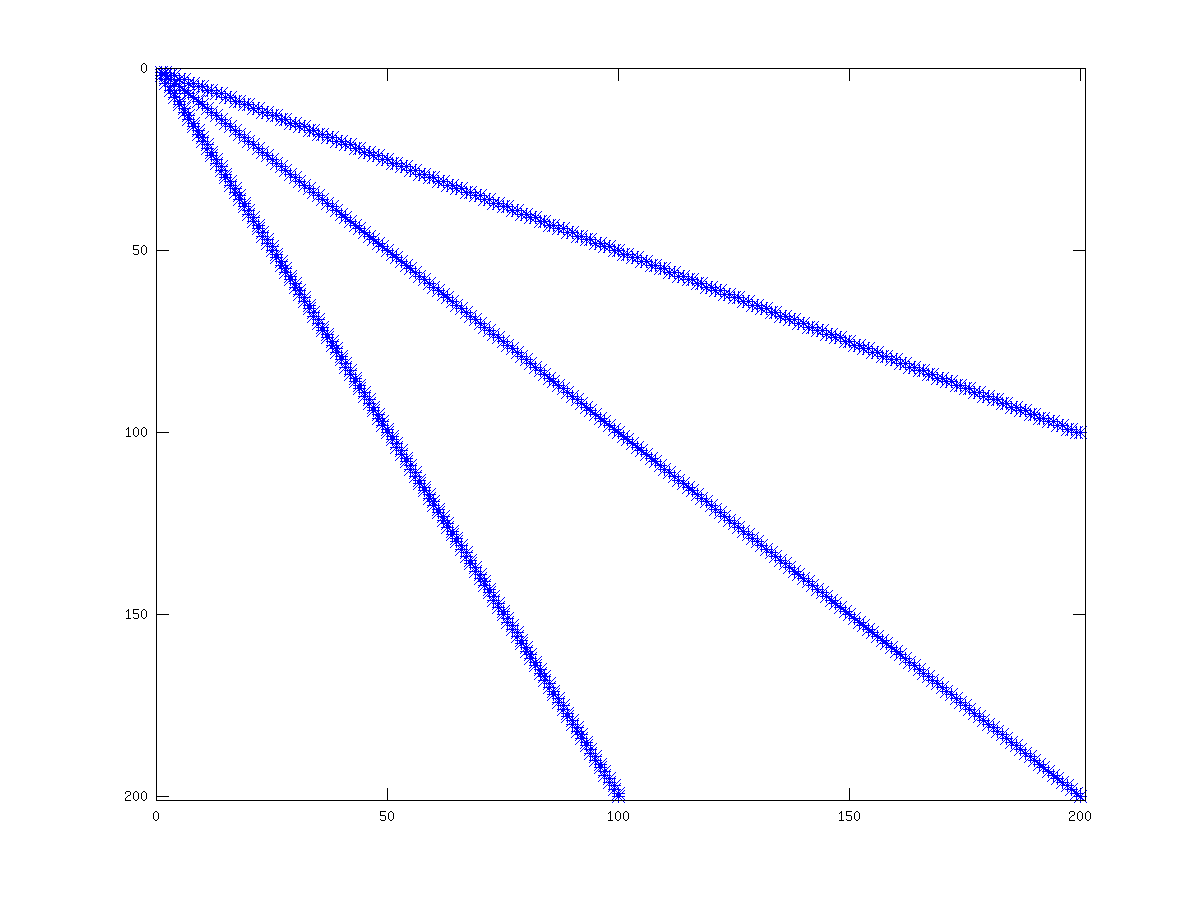
See fig:simplematrix, for an example of the structure of a simple positive definite matrix.
Figure 21.3: Structure of simple sparse matrix.
The standard Cholesky factorization of this matrix can be
obtained by the same command that would be used for a full
matrix. This can be visualized with the command
r = chol(A); spy(r);.
The original matrix had
598
non-zero terms, while this Cholesky factorization has
10200,
with only half of the symmetric matrix being stored. This
is a significant level of fill in, and although not an issue
for such a small test case, can represents a large overhead
in working with other sparse matrices.
The appropriate sparsity preserving permutation of the original
matrix is given by symamd and the factorization using this
reordering can be visualized using the command q = symamd(A);
r = chol(A(q,q)); spy(r). This gives
399
non-zero terms which is a significant improvement.
The Cholesky factorization itself can be used to determine the
appropriate sparsity preserving reordering of the matrix during the
factorization, In that case this might be obtained with three return
arguments as r[r, p, q] = chol(A); spy(r).
In the case of an asymmetric matrix, the appropriate sparsity
preserving permutation is colamd and the factorization using
this reordering can be visualized using the command q =
colamd(A); [l, u, p] = lu(A(:,q)); spy(l+u).
Finally, Octave implicitly reorders the matrix when using the div (/) and ldiv (\) operators, and so no the user does not need to explicitly reorder the matrix to maximize performance.
- Loadable Function: p = amd (s)
- Loadable Function: p = amd (s, opts)
Returns the approximate minimum degree permutation of a matrix. This permutation such that the Cholesky factorization of
s (p, p)tends to be sparser than the Cholesky factorization of s itself.amdis typically faster thansymamdbut serves a similar purpose.The optional parameter opts is a structure that controls the behavior of
amd. The fields of these structure are- opts.dense
Determines what
amdconsiders to be a dense row or column of the input matrix. Rows or columns with more thanmax(16, (dense * sqrt (n)entries, where n is the order of the matrix s, are ignored byamdduring the calculation of the permutation The value of dense must be a positive scalar and its default value is 10.0- opts.aggressive
If this value is a non zero scalar, then
amdperforms aggressive absorption. The default is not to perform aggressive absorption.
The author of the code itself is Timothy A. Davis (davis@cise.ufl.edu), University of Florida (see http://www.cise.ufl.edu/research/sparse/amd).
- Loadable Function: p = ccolamd (s)
- Loadable Function: p = ccolamd (s, knobs)
- Loadable Function: p = ccolamd (s, knobs, cmember)
- Loadable Function: [p, stats] = ccolamd (…)
Constrained column approximate minimum degree permutation.
p = ccolamd (s)returns the column approximate minimum degree permutation vector for the sparse matrix s. For a non-symmetric matrix s,s (:, p)tends to have sparser LU factors than s.chol (s (:, p)' * s (:, p))also tends to be sparser thanchol (s' * s).p = ccolamd (s, 1)optimizes the ordering forlu (s (:, p)). The ordering is followed by a column elimination tree post-ordering.knobs is an optional one- to five-element input vector, with a default value of
[0 10 10 1 0]if not present or empty. Entries not present are set to their defaults.-
knobs(1) if nonzero, the ordering is optimized for
lu (S (:, p)). It will be a poor ordering forchol (s (:, p)' * s (:, p)). This is the most important knob for ccolamd.-
knob(2) if s is m-by-n, rows with more than
max (16, knobs (2) * sqrt (n))entries are ignored.-
knob(3) columns with more than
max (16, knobs (3) * sqrt (min (m, n)))entries are ignored and ordered last in the output permutation (subject to the cmember constraints).-
knob(4) if nonzero, aggressive absorption is performed.
-
knob(5) if nonzero, statistics and knobs are printed.
cmember is an optional vector of length n. It defines the constraints on the column ordering. If
cmember (j) = c, then column j is in constraint set c (c must be in the range 1 to n). In the output permutation p, all columns in set 1 appear first, followed by all columns in set 2, and so on.cmember = ones(1,n)if not present or empty.ccolamd (s, [], 1 : n)returns1 : np = ccolamd (s)is about the same asp = colamd (s). knobs and its default values differ.colamdalways does aggressive absorption, and it finds an ordering suitable for bothlu (s (:, p))andchol (S (:, p)' * s (:, p)); it cannot optimize its ordering forlu (s (:, p))to the extent thatccolamd (s, 1)can.stats is an optional 20-element output vector that provides data about the ordering and the validity of the input matrix s. Ordering statistics are in
stats (1 : 3).stats (1)andstats (2)are the number of dense or empty rows and columns ignored by CCOLAMD andstats (3)is the number of garbage collections performed on the internal data structure used by CCOLAMD (roughly of size2.2 * nnz (s) + 4 * m + 7 * nintegers).stats (4 : 7)provide information if CCOLAMD was able to continue. The matrix is OK ifstats (4)is zero, or 1 if invalid.stats (5)is the rightmost column index that is unsorted or contains duplicate entries, or zero if no such column exists.stats (6)is the last seen duplicate or out-of-order row index in the column index given bystats (5), or zero if no such row index exists.stats (7)is the number of duplicate or out-of-order row indices.stats (8 : 20)is always zero in the current version of CCOLAMD (reserved for future use).The authors of the code itself are S. Larimore, T. Davis (Uni of Florida) and S. Rajamanickam in collaboration with J. Bilbert and E. Ng. Supported by the National Science Foundation (DMS-9504974, DMS-9803599, CCR-0203270), and a grant from Sandia National Lab. See http://www.cise.ufl.edu/research/sparse for ccolamd, csymamd, amd, colamd, symamd, and other related orderings.
-
- Loadable Function: p = colamd (s)
- Loadable Function: p = colamd (s, knobs)
- Loadable Function: [p, stats] = colamd (s)
- Loadable Function: [p, stats] = colamd (s, knobs)
Column approximate minimum degree permutation.
p = colamd (s)returns the column approximate minimum degree permutation vector for the sparse matrix s. For a non-symmetric matrix s,s (:,p)tends to have sparser LU factors than s. The Cholesky factorization ofs (:,p)' * s (:,p)also tends to be sparser than that ofs' * s.knobs is an optional one- to three-element input vector. If s is m-by-n, then rows with more than
max(16,knobs(1)*sqrt(n))entries are ignored. Columns with more thanmax(16,knobs(2)*sqrt(min(m,n)))entries are removed prior to ordering, and ordered last in the output permutation p. Only completely dense rows or columns are removed ifknobs (1)andknobs (2)are < 0, respectively. Ifknobs (3)is nonzero, stats and knobs are printed. The default isknobs = [10 10 0]. Note that knobs differs from earlier versions of colamdstats is an optional 20-element output vector that provides data about the ordering and the validity of the input matrix s. Ordering statistics are in
stats (1:3).stats (1)andstats (2)are the number of dense or empty rows and columns ignored by COLAMD andstats (3)is the number of garbage collections performed on the internal data structure used by COLAMD (roughly of size2.2 * nnz(s) + 4 * m + 7 * nintegers).Octave built-in functions are intended to generate valid sparse matrices, with no duplicate entries, with ascending row indices of the nonzeros in each column, with a non-negative number of entries in each column (!) and so on. If a matrix is invalid, then COLAMD may or may not be able to continue. If there are duplicate entries (a row index appears two or more times in the same column) or if the row indices in a column are out of order, then COLAMD can correct these errors by ignoring the duplicate entries and sorting each column of its internal copy of the matrix s (the input matrix s is not repaired, however). If a matrix is invalid in other ways then COLAMD cannot continue, an error message is printed, and no output arguments (p or stats) are returned. COLAMD is thus a simple way to check a sparse matrix to see if it's valid.
stats (4:7)provide information if COLAMD was able to continue. The matrix is OK ifstats (4)is zero, or 1 if invalid.stats (5)is the rightmost column index that is unsorted or contains duplicate entries, or zero if no such column exists.stats (6)is the last seen duplicate or out-of-order row index in the column index given bystats (5), or zero if no such row index exists.stats (7)is the number of duplicate or out-of-order row indices.stats (8:20)is always zero in the current version of COLAMD (reserved for future use).The ordering is followed by a column elimination tree post-ordering.
The authors of the code itself are Stefan I. Larimore and Timothy A. Davis (davis@cise.ufl.edu), University of Florida. The algorithm was developed in collaboration with John Gilbert, Xerox PARC, and Esmond Ng, Oak Ridge National Laboratory. (see http://www.cise.ufl.edu/research/sparse/colamd)
- Function File: p = colperm (s)
Returns the column permutations such that the columns of
s (:, p)are ordered in terms of increase number of non-zero elements. If s is symmetric, then p is chosen such thats (p, p)orders the rows and columns with increasing number of non zeros elements.
- Loadable Function: p = csymamd (s)
- Loadable Function: p = csymamd (s, knobs)
- Loadable Function: p = csymamd (s, knobs, cmember)
- Loadable Function: [p, stats] = csymamd (…)
For a symmetric positive definite matrix s, returns the permutation vector p such that
s(p,p)tends to have a sparser Cholesky factor than s. Sometimescsymamdworks well for symmetric indefinite matrices too. The matrix s is assumed to be symmetric; only the strictly lower triangular part is referenced. s must be square. The ordering is followed by an elimination tree post-ordering.knobs is an optional one- to three-element input vector, with a default value of
[10 1 0]if present or empty. Entries not present are set to their defaults.-
knobs(1) If s is n-by-n, then rows and columns with more than
max(16,knobs(1)*sqrt(n))entries are ignored, and ordered last in the output permutation (subject to the cmember constraints).-
knobs(2) If nonzero, aggressive absorption is performed.
-
knobs(3) If nonzero, statistics and knobs are printed.
cmember is an optional vector of length n. It defines the constraints on the ordering. If
cmember(j) = s, then row/column j is in constraint set c (c must be in the range 1 to n). In the output permutation p, rows/columns in set 1 appear first, followed by all rows/columns in set 2, and so on.cmember = ones(1,n)if not present or empty.csymamd(s,[],1:n)returns1:n.p = csymamd(s)is about the same asp = symamd(s). knobs and its default values differ.stats (4:7)provide information if CCOLAMD was able to continue. The matrix is OK ifstats (4)is zero, or 1 if invalid.stats (5)is the rightmost column index that is unsorted or contains duplicate entries, or zero if no such column exists.stats (6)is the last seen duplicate or out-of-order row index in the column index given bystats (5), or zero if no such row index exists.stats (7)is the number of duplicate or out-of-order row indices.stats (8:20)is always zero in the current version of CCOLAMD (reserved for future use).The authors of the code itself are S. Larimore, T. Davis (Uni of Florida) and S. Rajamanickam in collaboration with J. Bilbert and E. Ng. Supported by the National Science Foundation (DMS-9504974, DMS-9803599, CCR-0203270), and a grant from Sandia National Lab. See http://www.cise.ufl.edu/research/sparse for ccolamd, csymamd, amd, colamd, symamd, and other related orderings.
-
- Loadable Function: p = dmperm (s)
- Loadable Function: [p, q, r, s] = dmperm (s)
-
Perform a Dulmage-Mendelsohn permutation on the sparse matrix s. With a single output argument dmperm performs the row permutations p such that
s (p,:)has no zero elements on the diagonal.Called with two or more output arguments, returns the row and column permutations, such that
s (p, q)is in block triangular form. The values of r and s define the boundaries of the blocks. If s is square thenr == s.The method used is described in: A. Pothen & C.-J. Fan. Computing the block triangular form of a sparse matrix. ACM Trans. Math. Software, 16(4):303-324, 1990.
- Loadable Function: p = symamd (s)
- Loadable Function: p = symamd (s, knobs)
- Loadable Function: [p, stats] = symamd (s)
- Loadable Function: [p, stats] = symamd (s, knobs)
For a symmetric positive definite matrix s, returns the permutation vector p such that
s (p, p)tends to have a sparser Cholesky factor than s. Sometimes SYMAMD works well for symmetric indefinite matrices too. The matrix s is assumed to be symmetric; only the strictly lower triangular part is referenced. s must be square.knobs is an optional one- to two-element input vector. If s is n-by-n, then rows and columns with more than
max(16,knobs(1)*sqrt(n))entries are removed prior to ordering, and ordered last in the output permutation p. No rows/columns are removed ifknobs(1) < 0. Ifknobs (2)is nonzero,statsand knobs are printed. The default isknobs = [10 0]. Note that knobs differs from earlier versions of symamd.stats is an optional 20-element output vector that provides data about the ordering and the validity of the input matrix s. Ordering statistics are in
stats (1:3).stats (1) = stats (2)is the number of dense or empty rows and columns ignored by SYMAMD andstats (3)is the number of garbage collections performed on the internal data structure used by SYMAMD (roughly of size8.4 * nnz (tril (s, -1)) + 9 * nintegers).Octave built-in functions are intended to generate valid sparse matrices, with no duplicate entries, with ascending row indices of the nonzeros in each column, with a non-negative number of entries in each column (!) and so on. If a matrix is invalid, then SYMAMD may or may not be able to continue. If there are duplicate entries (a row index appears two or more times in the same column) or if the row indices in a column are out of order, then SYMAMD can correct these errors by ignoring the duplicate entries and sorting each column of its internal copy of the matrix S (the input matrix S is not repaired, however). If a matrix is invalid in other ways then SYMAMD cannot continue, an error message is printed, and no output arguments (p or stats) are returned. SYMAMD is thus a simple way to check a sparse matrix to see if it's valid.
stats (4:7)provide information if SYMAMD was able to continue. The matrix is OK ifstats (4)is zero, or 1 if invalid.stats (5)is the rightmost column index that is unsorted or contains duplicate entries, or zero if no such column exists.stats (6)is the last seen duplicate or out-of-order row index in the column index given bystats (5), or zero if no such row index exists.stats (7)is the number of duplicate or out-of-order row indices.stats (8:20)is always zero in the current version of SYMAMD (reserved for future use).The ordering is followed by a column elimination tree post-ordering.
The authors of the code itself are Stefan I. Larimore and Timothy A. Davis (davis@cise.ufl.edu), University of Florida. The algorithm was developed in collaboration with John Gilbert, Xerox PARC, and Esmond Ng, Oak Ridge National Laboratory. (see http://www.cise.ufl.edu/research/sparse/colamd)
- Loadable Function: p = symrcm (S)
Symmetric reverse Cuthill-McKee permutation of S. Return a permutation vector p such that
S (p, p)tends to have its diagonal elements closer to the diagonal than S. This is a good preordering for LU or Cholesky factorization of matrices that come from 'long, skinny' problems. It works for both symmetric and asymmetric S.The algorithm represents a heuristic approach to the NP-complete bandwidth minimization problem. The implementation is based in the descriptions found in
E. Cuthill, J. McKee: Reducing the Bandwidth of Sparse Symmetric Matrices. Proceedings of the 24th ACM National Conference, 157–172 1969, Brandon Press, New Jersey.
Alan George, Joseph W. H. Liu: Computer Solution of Large Sparse Positive Definite Systems, Prentice Hall Series in Computational Mathematics, ISBN 0-13-165274-5, 1981.
| [ < ] | [ > ] | [ << ] | [ Up ] | [ >> ] | [Top] | [Contents] | [Index] | [ ? ] |